Dear Brew enthusiasts,
I know it has been a little while, but due to a much-needed break (holiday) and travel (within the country), I have not sent you any newsletter posts—apologies for that.
I am happy to tell you that I am now back refreshed (and double vaccinated) to bring you the latest beer brewing research. This week, we will be talking about some exciting work on beer haziness or turbidity. Specifically, I will describe the identification of proteins that may be responsible for unwanted haziness in beer and can tell us about problems with our brew. Understanding which proteins cause or are associated with hazy beer, can help us identify the actual problem.
As many of you will appreciate, beer haziness is not necessarily an issue. Some beer styles (e.g. wheat beers) are inherently turbid, and most beer lovers do not mind sampling other somewhat cloudy brews. But what if you are a brewery producing a beer that features an unwanted haze? The only reasonable way to fix such a problem is to identify it through troubleshooting. The work I am describing below does just that.

A group of researchers, led by Dr Dawn Maskell, investigated the formation of a haze in a commercially available India Pale Ale. Initial investigation (not described here) excluded microbial contamination as the culprit (using size-based filtration), leading to the suggestion that another component formed the problem. One of the hypotheses put forward was that haze formation was due to elevated levels of proteins responsible for hazing. The authors (Huismann et al) set out to test this hypothesis by investigating the protein composition in high haze and low haze beer samples. You can find their research here.
The researchers sampled high haze and low haze beers and isolated all the proteins from these samples. To do so, the researchers mixed their samples with trichloroacetic acid (TCA), a compound that, when combined with proteinaceous (cold) solutions, make proteins clump together into heavier particles. By centrifuging such samples, you can apply high gravity levels to make these particles drop to the bottom of samples (pellet), which can then be washed, cleaned and dissolved again in buffers that allow their analyses.
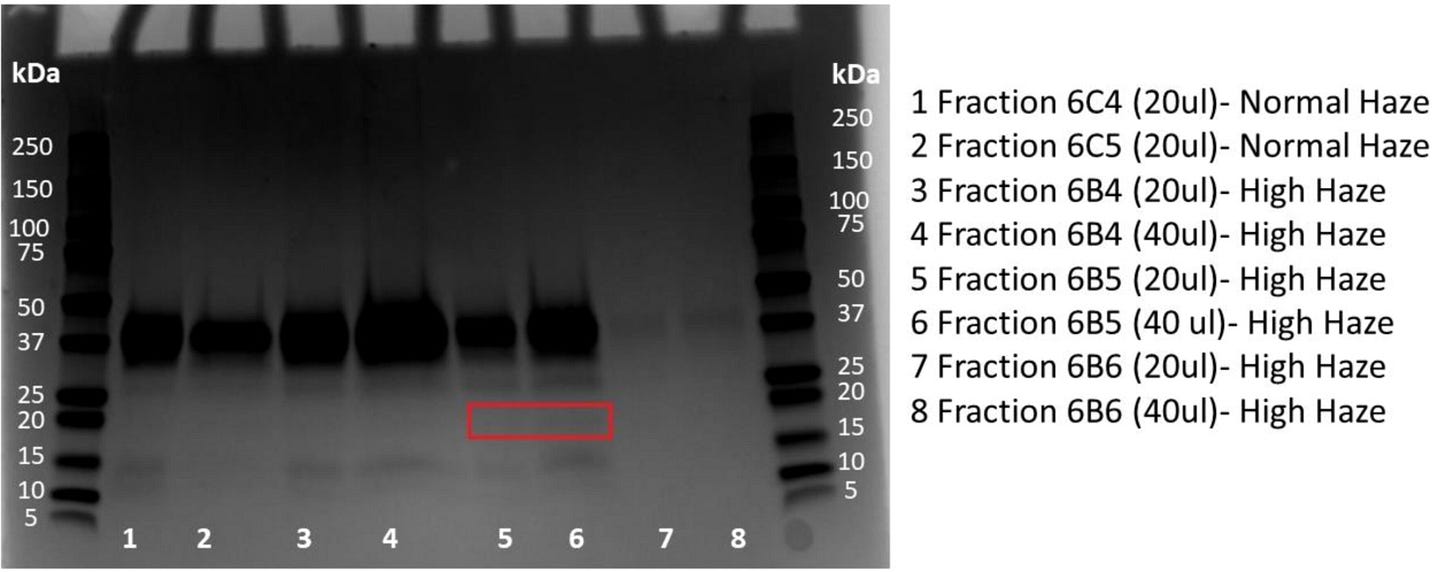
Having isolated most proteins from each sample, the authors then fractionated their protein samples and separated them onto special gels (SDS-PAGE gels). By applying a stain that can visualise proteins that have migrated through the gel (I won't go into detail here about the technique), the authors could visualise specific protein bands that appear to be specific to high-haze beer samples. The implication, therefore, is that (i) these proteins either are directly responsible for beer hazing or (ii) give important clues about the origin of the problem (e.g. is the haze coming from grains or yeast?).
Therefore, the authors set out to identify the proteins that appear elevated in the high haze samples. To find such proteins, the authors used a sophisticated analytical technique called LC-QTOF-MS. This method uses liquid chromatography (LC) to separate proteins (or their digested counterparts) into fractions that get injected into a mass spectrometer (MS) that measures the time of flight (TOF) of protein fragments.
As each protein or peptide has its own unique TOF profile (depending on size, sequence and charge), we can determine their sequences by matching them across established databases. Knowing each peptide sequence and counting the number of peptides that match each given protein, we can estimate the number of protein molecules for a given candidate. By comparing the number of "hits" or the level of coverage of a protein between samples, we can measure and estimate differences in the abundance of a protein between samples. Here, this means between high and low haze beer samples.
In this way, the authors made several interesting observations. Firstly, they identified a fair number of mannoproteins. These are proteins that sit below the yeast cell wall and help achieve cell wall integrity, rigidity and cell shape. Since they sit inside the yeast cell, their specific presence in high haze beer samples suggests their release from (dead) yeast.
Another exciting observation concerned another class of yeast proteins, called the flocculation proteins. These proteins were far more abundant in the high haze beer samples when compared to the low haze controls. Given that yeast also produces flocculation proteins, the problem of haze formation in the commercial IPA likely originates with yeast. The observation that both mannoproteins and flocculation proteins are released when yeast cells are stressed, gives further weight to this hypothesis.
What does this all mean, and why is it important?
This work is not novel from a purely scientific point of view (and on a fundamental biology level). Other studies have also investigated the presence and identity of haze proteins in beer. Consequently, many such proteins are now known. However, the exciting aspect of this work is that by using proteomic analyses of problematic beer samples, researchers (and brewers) can detect biological signatures that signal the nature of the problem with your beer or brew process. In other words, proteomics may emerge as an important analytical tool that can help us diagnose issues or even assess beer quality.
I think that this is a significant advance worth mulling over whilst enjoying your next (hazy) brew.
Have a great weekend!
Cheers!
Edgar, The Beerologist.



